SBU’s Balázsi’s work has implications for drug resistance in cancer
By Daniel Dunaief
Take two identical twins with the same builds, skill sets and determination. One of them may become a multimillionaire, a household name and the face of a franchise, while the other may toil away at the sport for a few years until deciding to pursue other interests.
What causes the paths of these two potential megastars to diverge?
Gábor Balázsi, an associate professor in biomedical engineering at Stony Brook University, asked a similar question about a cellular circuit in the hopes of learning more about cancer. He wanted to know what is it about the heterogeneity of a cancer cell that makes one susceptible to treatment from chemotherapeutic drugs and the other resistant to them. Heterogeneity comes from molecular differences where the original causes may be subtle, such as two molecules colliding or a cell being closer to the tumor’s surface, while the consequences can create significant differences, even among cells with the same genes.
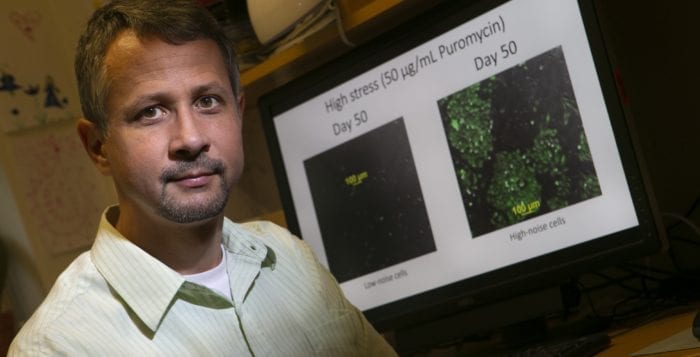
In research published this week in the journal Nature Communications, Balázsi used two mammalian cell lines that were identical except that each carried a different synthetic gene circuit that made one more heterogeneous than the other. He subjected the two cell lines, which would otherwise perform the same function, to various levels of the same drug to determine what might cause one to be treatable and the other to become resistant.
Through these mammalian cells, Balázsi created two circuits, one of which kept the differences between the cells low, while the other caused larger differences. Once inserted in the cell, these gene circuits created uniform and variable populations that could serve as models for low and high heterogeneity in cancer.
Working with Kevin Farquhar, who recently graduated from Balázsi’s lab, and former Stony Brook postdoc Daniel Charlebois, who is currently at the Department of Physics at the University of Alberta, Balázsi tried to test how uniform versus heterogeneous cell populations respond to treatment with different drug levels.
Using the two synthetic gene circuits in separate but identical cell lines, the Stony Brook scientists, with financial support from the National Institutes of Health and the Laufer Center for Physical and Quantitative Biology at SBU, could re-create high and low stochasticity, or noise, in drug resistance in two cell lines that were otherwise identical.
While the work is in its preliminary stages and is a long way from the complicated collection of genes responsible for various types of cancer, this kind of analysis can test the importance of specific processes for drug resistance.
“Only in the last decade or so have we come to realize how much heterogeneity (genetic and nongenetic differences) can exist within a tumor in a single patient,” Patricia Thompson-Carino, a professor in the Department of Pathology at the Renaissance School of Medicine at SBU, explained in an email. “Thinking of cancer in a single patient as several different diseases is a bit daunting, though currently, this heterogeneity and its direct effects on how the cancer behaves remains poorly understood.”
Indeed, Thompson-Carino added that she believes Balázsi’s work will “shed light on cancer cell responses to therapy. With the rise in cancer therapies designed to specific targets and the resistance that emerges in patients on these therapies, I think [Balázsi’s] work is of extremely high value” which may help with the puzzle of how nongenetic or epigenetic heterogeneity affects responses to treatment, she continued.
In the future, researchers and clinicians may look to develop new ways of biomarker analysis that considers the variability, rather than just the average level of a biomarker.
Balázsi suggested that looking only at the variability of cells is analogous to observing an iron block sinking in water. Someone might conclude that all solids sink in liquids. Similarly, scientists might decide that cellular variability always promotes drug resistance from observations when this happens. To gain a fuller understanding of the effect of variability, however, researchers need to equalize the averages. They then need to explore what happens at various levels of drug treatment.
Current therapies do not target heterogeneity. If such future treatments existed, doctors and scientists could combine ways of treating heterogeneity with attacking cancer, which might work in the short term or prevent cancer from recurring.
Balázsi suggests his paper is a part of his attempt to address three different areas. First, he’d like to figure out how to categorize patients better, including the variability of biomarkers. Second, he believes this kind of analysis will assist in creating future combinations of treatments. By understanding how the variability of cancer cells contributes to its reaction to therapies, he might help create a cocktail of treatments, akin to the effort that helped with the treatment of HIV in the lab.
Third, he’d like to obtain cancer samples and allow them to evolve in a lab, where he can check to see how they respond to treatment levels and administration scheduling. This effort could allow him to determine the optimal drug combination and dosing for a patient.
For the work that led to the current Nature Communications paper, Balázsi explored how mammalian cells respond to various concentrations of a drug. Over 80 percent of the genes in these cells are also present in human cells, so the mechanisms he discovered and conclusions he draws should apply to human cancer cells as well.
He concluded that cells with more heterogeneity, where the cells deviate more from the average, resist drugs better when the drug level is high. These same cells, show greater sensitivity when the drug is low.
Balázsi recognizes that the work he’s exploring is a “complex problem” and that it requires considerable additional research to understand and appreciate how a therapy might kill one cancer cell, while the same treatment in the same environment doesn’t have the same effect on a genetically identical cell.