By Daniel Dunaief
The raw materials were here. Somehow, billions of years ago, these materials followed patterns and repeated and revised the process, turning the parts into something more than a primordial soup.
Ken Dill, who is a distinguished professor and the director of the Laufer Center for Physical and Quantitative Biology at Stony Brook University, took a methodical approach to this fundamental development. He wanted to understand the early statistical mechanics that would allow molecules to form long chains, called polymers, which contained information worthy of being passed along. The process of forming these chains had to be self-sustaining.
After all, Dill said, many activities reach an end point. Putting salt in water, for example, creates a mixture, until it stops. Dill, however, was looking for a way to understand auto-catalytic or runaway events. Lighting a forest fire, for example, is much more self sustaining, although even it eventually stops. Life has continued for over four billion years.
On Aug. 22, Dill, Elizaveta Guseva and Ronald Zuckermann, the facility director in biological nanostructures at the Lawrence Berkeley National Laboratory, published a paper in the journal Proceedings of the National Academy of Sciences (PNAS).
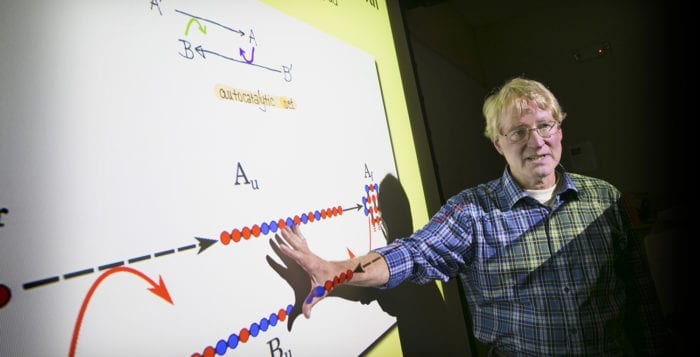
The researchers developed a fold and catalyze computational model that would explain how these long chains developed in a self-sustaining way, in which hydrophilic and hydrophobic polymers fold and bind together.
Random sequence chains of each type can collapse and fold into structures that expose their hydrophobic parts. Like a conga line at a wedding reception, the parts can then couple together to form longer chains.
These random chemical processes could lead to pre-proteins. Today’s proteins, Dill said, mostly fold into a very particular shape. Pre-proteins would have been looser, with more shape shifting.
The workhorses of the body, proteins perform thousands of biochemical reactions. Dill suggested that this model “rates high on the list” in terms of the findings he’s made over the course of his career.
Zuckermann described this work as significant because it lays out predictions that can be tested. It highlights the importance of chemical sequence information in polymer chains and “how certain sequences are more likely to fold into enzyme-like shapes and act as catalysts than others,” he explained in an email.
Zuckermann works with substances he figured out how to make in a lab that are called peptoids, which are non-natural polymers. These peptoids are a “good system to test the universality of [Dill’s] predictions,” he said.
The “beauty” of Dill’s work, Zuckermann suggested, is that “it should apply to most any kind of polymer system” where researchers control the monomer sequence and include hydrophobic and hydrophilic monomers in a particular order, putting Dill’s predictions to the test.
For her part, Guseva worked in Dill’s lab for her PhD thesis. She had started her research on something that was “more standard physical biology” Dill said, but it “was not turning out to be particularly interesting.”
The scientists had a discussion about trying to develop a chemical model related to the origins of life. While exciting for the scope of the question, the research could have come up empty.
“There was so much potential to fail,” Dill said. “I feel pretty uncomfortable in general about asking a graduate student to go in that direction, but she was fearless.”
Dill and Zuckermann, who have collaborated for over 25 years, are trying to move forward to the next set of questions.
Zuckermann’s efforts will focus on finding catalytic peptoid sequences, which are nonbiological polymers. He will synthesize tens of thousands of peptoid sequences and rank them on how enzyme-like they are. This, he explained, will lead to a better understanding of which monomer sequences encode for protein-like structure and function.
Zuckermann suggested that the process in this research could have the effect of transforming a soup of monomers into a soup of functional polymers. This, he said, might set the stage for the evolution of DNA and RNA.
Proteins could have been a first step towards a genetic code, although life, as currently defined, would not have blossomed until a genetic code occurred, too, Dill suggested.
The origins of DNA, however, remains an unanswered question. “We’re trying to think about where the genetic code comes from,” Dill said. “It’s not built into our model per se. Why would biology want to do a two polymer solution, which is messy and complicated and why are proteins the functional molecules? This paper doesn’t answer that question.”
Dill and Zuckermann are in the early stage of exploring that question and Dill is hopeful he can get to a new model, although he doesn’t have it yet.
Dill moved from the University of California at San Francisco to join the Laufer Center about seven years ago. He appreciates the freedom to ask “blue sky questions” that he couldn’t address as much in his previous work.

A resident of Port Jefferson, Dill lives with his wife Jolanda Schreurs, who has a PhD in pharmacology. The couple has two sons, Tyler and Ryan.
Tyler graduated with a PhD from the University of California at San Diego and now works for Illumina, a company which which makes DNA sequencers. Ryan, meanwhile, is earning his PhD in chemistry from the University of Colorado and is working on lasers.
“We didn’t try to drag our sons into science,” Dill said. “With both kids, however, we had a workshop in the basement” where they often took anything that was within arm’s reach and nailed it to a board. One of the finished products was a remote-controlled and motorized boat.
As for his lab work, Dill is thrilled to have this model that he, Guseva and Zuckermann provided, while he recognizes the questions ahead. Scientists “see something puzzling and, rather than saying, ‘I need to avoid this, I don’t have an answer,’ we find it intriguing and these things lead from one step to the next. There tends to remain a huge number of super fascinating problems.”